Research
Science’s ‘Mother of Ribbon Diagrams’ celebrates 50 years at Duke
Science’s ‘Mother of Ribbon Diagrams’ celebrates 50 years at Duke
by Lindsay Key
In almost every scientific paper that describes proteins you’ll find Duke biochemist Jane Richardson’s handiwork. The images, commonly referred to as “ribbon diagrams,” are basic to the language of protein science. Ranging in likeness from a flapping kite tail to a tight coil of crimped gift ribbon, these carefully etched diagrams invented by Richardson have served for many years as a primary way that scientists can describe what they see in their data.
Often called the ‘building blocks of life,’ proteins are themselves a sort of language. Found in every living organism, their 3D shapes determine the function and structure of body tissues such as muscle or skin and chemical functions such as breathing, digestion or growth. Understanding their precise structure and interactions is important for drug design to improve human health. Some of the most widely-used drugs, including the antibiotic penicillin, have been redesigned and improved due to better understanding of the proteins they target.
Today, advances in imaging technology, such as cryo-electron microscopy and free-electron lasers, allow scientists to freeze protein samples to image huge, dynamic molecular structures or to collect the information from instantaneous “diffract and destroy” of tiny droplet samples using electron blasts. But in the 1960s and 1970s, it usually took scientists many years to solve just one small structure.
The great protein race
Jane Richardson and her husband, biochemist Dave Richardson, came to Duke in 1969. At the time, in their late twenties, they had already had illustrious scientific careers. Meeting as undergraduates at Swarthmore College in Pennsylvania, they migrated to Boston together, where Jane completed a master’s degree in philosophy at Harvard University, and Dave completed a Ph.D. in chemistry at MIT.
After her graduation, Jane joined Dave in the MIT lab of Al Cotton, where they spent seven years solving the structure of Staphylococcal nuclease, an enzyme that cleaves DNA and RNA. Enzymes are proteins that are folded into complex shapes that allow smaller molecules to fit into them and increase the rate of chemical reactions in the body.
When the Richardsons arrived at Duke, they were quickly pulled into work with other biochemistry department researchers on studying superoxide dismutase, an enzyme that protects all living things against the toxicity of oxygen. The protein's function had just been identified by Duke scientists Irwin Fridovich and his graduate student Joe McCord, and the Richardsons solved its 3-dimensional structure using X-ray crystallography. The technique involves placing a molecular crystal in a beam of X-ray light, and then measuring the ways that the light diffracts from the crystal in order to get a three-dimensional picture of the density of electrons, and thus atoms, in the molecule.

A photo of Jane and Dave Richardson from 1971 pinned on a board in their lab at Duke.
At that time, only about twenty different protein structures had been solved in the world, and the Richardsons did two of them. Their field, called structural biology, gained momentum as scientists raced to solve as many molecular shapes as possible. The Richardsons began to look for patterns in the small collection of already determined protein structures.
Jane Richardson's ribbon drawing of Cu,Zn superoxide dismutase (PDB file 1sod), ink outline with colored pencil.
One of their biggest discoveries came through the doorway — literally — when they ran into NIH molecular biologist David Davies at a regional crystallography meeting. Davies was carrying a wire model of immunoglobin which looked eerily similar to the Richardsons’ own wire model of superoxide dismutase.
Sitting down to compare the structures closely, the Richardsons discovered that both structures had the same crisscrossing of beta strands, a pattern they named the ‘Greek key fold’ due to its similarity to a design common on vases, jewelry, and placemats. This helped them to realize that pattern similarities among proteins can be due to folding preferences. The discovery was key to understanding the underlying rules of protein structure.
Deciding what’s important
As the elegance and complexity of proteins became more evident, scientists needed something better than wire models to articulate the subtleties they were identifying, especially in print journals. Several people had published schematic drawings of individual proteins, but each used different conventions and viewpoints. For a major review article comparing folds and details of all the known structures, Jane Richardson needed to develop a good, consistent system. She picked up a pencil and began to try drawing smooth ribbons over the atoms in early, crude computer-graphics printouts. It was promising, but hard to correctly convey the proteins in three dimensions.
“I’m not an artist and I’m not very good at drawing anything else,” she said. “It took a very long time to figure this out, how you can really show the 3D relationships on a 2D page, and also keep them accurate to the experimental atom coordinates.”
Richardson interviewed her mother-in-law, who was an artist, and reached out to Irving Geis, the artist behind early Scientific American illustrations, for advice on how to complete the drawings. But in the end, she found that a belt and a solid philosophy education were most helpful.
She held up a twisted belt and studied what her two eyes actually saw of the edge and sides in order to learn how to draw twisted protein beta strands convincingly. She also relied on theories of illusion that she had learned in her psychology class at Swarthmore, such as how to slightly offset objects that cross at a low angle to make them look right.
“Figuring this out and trying out a lot of different things took probably half a year,” Richardson said. “Then, each drawing took from part of a day to a week depending on how hard it was to draw. Each time, you have to decide what’s important [to depict]. Then another year to write the 169 pages of text around those drawings.”
Jane Richardson's ribbon drawing of triose phosphate isomerase (PDB file 1tim), ink outline colored with pastels. Wikipedia picture of the day Nov. 19, 2009.
In 1981, Jane Richardson published her drawings for the first time in the journal Advances in Protein Chemistry. Scientists immediately started using them and some even argued with her about how she’d depicted a particular molecule, opening up a new era of debate about scientific interpretation. In 1985, she was awarded a MacArthur Fellowship, the so-called “genius grant.”
Dave Richardson remembers the time well. “It was a heady thing to see science be able to be built on illustrations,” he said. “The drawings were helping us to set how we think about structures, and how we influence each other.”
Today, Jane’s ‘ribbon diagrams’ are used almost universally for overall description of proteins. Although they are now created using computer graphics rather than pen and ink, the language she invented remains the same. Those who remember where it came from still call it a “Richardson diagram.”
“It’s very satisfying that the graphics have come back around to look a lot like the drawings,” said Jane Richardson. “Not exactly, but similar. However, it’s also a little scary because, with the drawings, everybody realized they were subjective. But when you have computer graphics, they don’t realize that the programmer, or whoever made the algorithm for it, is making these choices. They’re deciding how you should look at the structure.”
Former student Gary Kapral, who graduated in 2013 with his Ph.D. from the Richardson lab, was inspired by the level of accuracy that Jane Richardson promoted in both her science and her art.
“Working with Jane was like working with da Vinci or Raphael—you knew that you would produce a work of art as well as scientific progress,” said Kapral. “Our publications and software were full of puns and allusions to great literary works hidden among the requisite references to the scientific giants on whose shoulders we stood. Our published figures were scrupulously designed to contain layers of information yet be easy to follow. No work environment I have experienced has come close to approaching the creativity and sheer joy of working in the Richardson lab.”
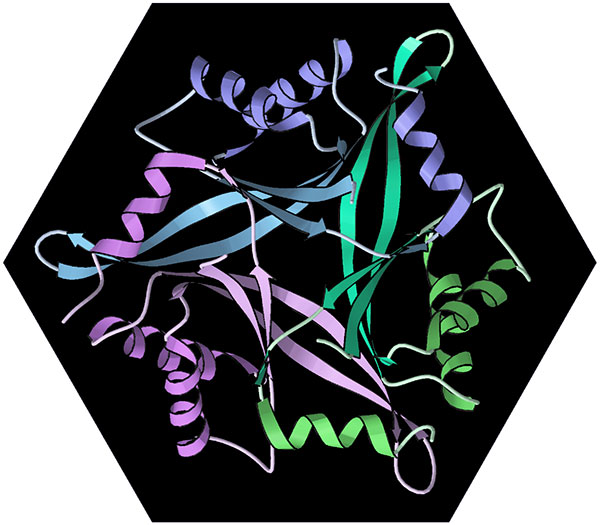
Computer ribbon diagram of the intertwined trimer of human CUTA protein (PDB file 1xk8), drawn in Dave Richardson’s MAGE graphics program.
Jane Richardson's ribbon drawing of Cu,Zn superoxide dismutase, again, superimposed with a rendering of the ribbons using the KiNG (Kinemage, Next Generation) software developed in the Richardson Laboratory.
Getting the perspective
When Jane Richardson thinks back to her educational choices, she wouldn’t change a thing. Unlike most biochemists, Richardson does not hold a Ph.D. in any science. Starting out as a math, physics, and astronomy major at Swarthmore, she switched to philosophy (with minors in math and physics) halfway through because she found the class discussions most interesting.
“The philosophy has been enormously useful,” she said. “A liberal arts education is the right way to get into science, I think. And then later, in graduate school, you can learn the technical stuff.” Jane Richardson learned biochemistry and biophysics working as a technician in her husband’s lab at MIT after receiving her master’s degree.
Dave Richardson said his wife’s philosophy background has benefited their shared lab in countless ways. “There is an impression that scientists think and question things, but as a matter of fact, they’re not better than anyone else,” he said. “The philosophers really question. For them, it’s not a matter of asking questions, but deciding what questions to ask. Jane is amazing at that—she can hold off for a long time before she jumps to any conclusions and I think that’s a real gift.”
Not only has Jane Richardson been trained to think critically, but she also makes a point to get away and make time for reflection. Every summer, the Richardsons disappear to their cabin high in the Sierra Nevada mountains, just a few miles off the John Muir Trail, to think. They backpack for a week, or even a month, winding through rocky back country trails as well as enjoying the spectacular view from their front porch.
They follow the advice of their long-ago mentor, biochemist Fred Richards, who made a point to get away during the summer. While Richards chose to sail up and down the Atlantic coast, the Richardsons backpack in the mountains every summer.
“A lot of the reason we do this, as Fred did, is that things are very competitive and time-conscious, and much more now than they were,” said Jane Richardson. “You’re always behind. And it always is that something should have been done last week or last year or whatever. So you keep pushing at whatever you’re doing without really having the time to step back and figure out if you should be doing something different or in a different way. We find that it takes us about a week to get to the stage where we’re away from things long enough to get that perspective. We have switched what we were doing a number of times and it mostly was ideas that we got while we were in the middle of a backpacking trip. Those new directions have included developing kinemage graphics, going into early protein de novo design, and having our first child. In contrast, it was one of our students, Laura Murray, who got us into studying the enlighteningly distinct but similar RNA molecules.”
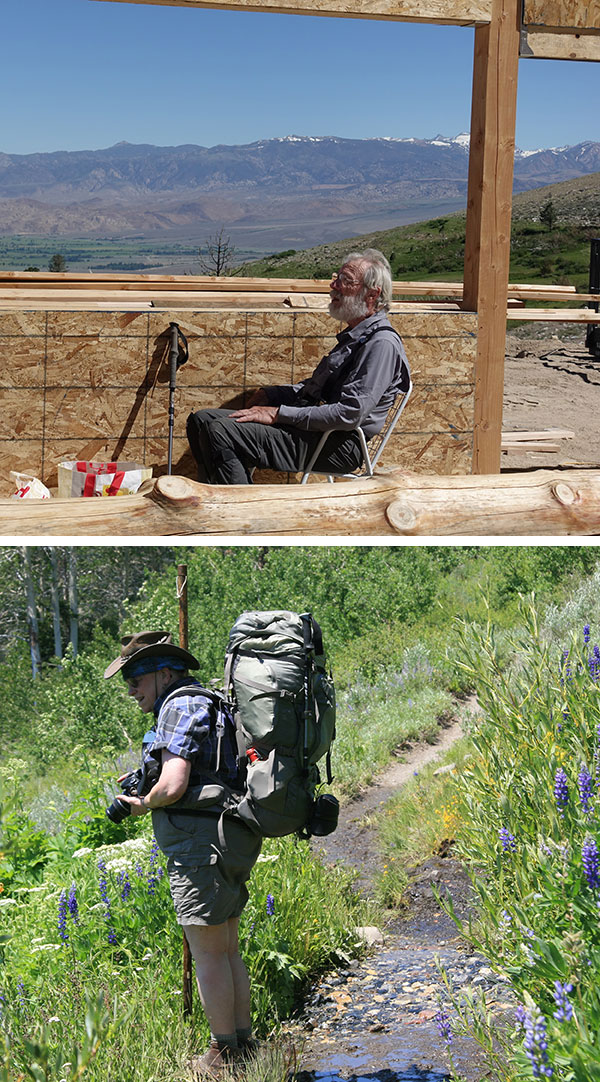

(top) Dave Richardson in the rebuild of their Sierra Nevada cabin after a wildfire. (bottom) Jane Richardson taking flower photos on a Sierra backpacking trip.
Forming the bond
The Richardsons have built so much together over the years — a career, a family (a son and a daughter) and three houses — two in Durham and one in California. Their current residence in Durham is a geodesic dome and an octagonal treehouse surrounded by the Korstian Division of Duke Forest and built to mimic a T-even virus — a shape they recognized in their build kit halfway through and made a point to fulfill.
One of Dave Richardson’s favorite things about being with Jane is that he doesn’t have to take long to explain himself — she just gets it. Both are visual thinkers and can communicate efficiently with each other about highly technical material.
“Jane and I can walk along a mountain trail talking about the intricate details inside a protein and know that we’re actually communicating, that we’re seeing the same thing in our mind’s eye,” said Dave Richardson.
Beyond that, they both have a passion for communicating with others in the scientific community, in order to build on the science that is available and make it better. They have many collaborations within and across departments at Duke, and also work with many international collaborative groups such as the Validation Task Force committees, the RNA Ontology Consortium, the CASP design and CryoEM Model Challenge competitions, and the Phenix structural-biology software project.
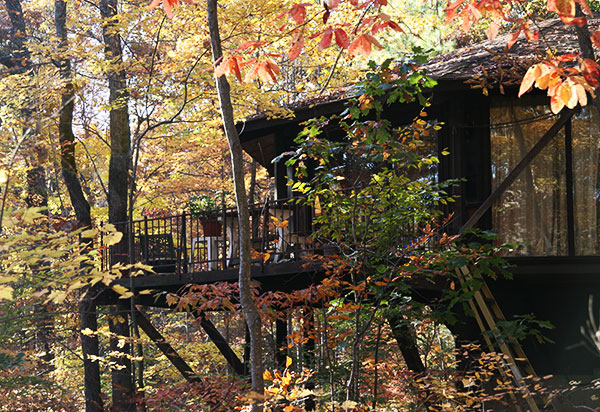

The Richardson's house in Duke Forest, designed to mimic the shape a T-even bacteriophage.
is a website they developed to help other scientists diagnose and correct local problems in their structures. Hydrogen atoms are not seen directly by protein crystallography, but the Richardsons realized they are crucial for understanding molecular contacts. Molprobity adds H atoms using chemical knowledge plus optimizing their hydrogen bonds and steric clashes, and then uses them to sensitively find problems in experimental structures. Once seen, those problems can usually be corrected, and the Richardsons' tools have helped both scientists and software to improve the structures deposited into the worldwide Protein DataBank.
Another rendering of the monomeric Cu,Zn superoxide dismutase using KiNG with additional detail and alternate display options creates a much different picture of the protein.
Now aged 77, Jane Richardson regularly contributes scientific images (and mountain and flower images) for public use to Wikimedia Commons. As president of the Biophysical Society, she started a WikiProject on Biophysics to encourage society members and others to edit and improve biophysics-related Wikipedia articles.
“Over the years, we have done very well by emphasizing collaboration over competition, talking freely about results, and encouraging our students to carry their projects away with them,” they write in their autobiography, published in 2013 in the Annual Review of Biophysics. “After all, there are many more great directions to follow up than we can possibly do ourselves, and science is about the joint building of an actively evolving network of information.”
Despite their shared passions and interests, the couple is also different in many ways. Jane has a knack for understanding complex mathematical concepts, while Dave is especially skilled in working with machinery — a talent that has served the lab well in recent years, in developing software as a machinery for understanding protein structure.
“When thinking of research done in the Richardson lab, it’s impossible to separate Dave and Jane,” said former student Bradley Hintze, who graduated with his Ph.D. from the lab in 2015. “They are two people but truly represent one research-investigator entity. Dave has the ability to dig deep in the methodology and provide detailed understanding down in the weeds. While Jane certainly has this ability, her real strength comes from her phenomenal ability to see the big picture. This allows her to see connections between disparate problems that the rest of us fail to see because we’re perceiving just the details.”
Inspiration and Support in the Richardson Lab
Colleagues in the biochemistry department at Duke are grateful to work in such close proximity and collaboration with the Richardsons.
“As structural biologists we know that macromolecules such as proteins have very complex structures,” said colleague Maria Schumacher, Ph.D. the Nanaline H. Duke Professor of Biochemistry. “They can be very difficult to understand when looking at the entire 3-D fold with all the atoms. What Jane and Dave developed was a standard (and very beautiful) way to visualize macromolecules and how they fold that enabled everyone to better understand their function. Very importantly, this is true not only for structural biologists, but for all scientists in diverse fields. As a result, Jane has had a profound impact in how we in science understand life at the molecular level.”
“Jane has been an inspiration to all of us here in Duke Biochemistry and to many other biochemists throughout the world,” said department chair Richard G. Brennan, Ph.D. “Her passion for understanding how biology works at the level of macromolecular structure and her outstanding contributions to the field of structural biology are unparalleled. Further, this zeal is carried over to the students and postdoctoral fellows, who have been lucky enough to be members of her laboratory or to have taken her structural biology course, which is co-taught by Dave.”
“Jane Richardson is one of the great structural biologists of our time,” said Paul Modrich, Ph.D., the James B. Duke Professor of Biochemistry and 2015 Nobel Laureate in chemistry. “Her elegant ribbon diagrams have become the standard for visualization of protein structures.”
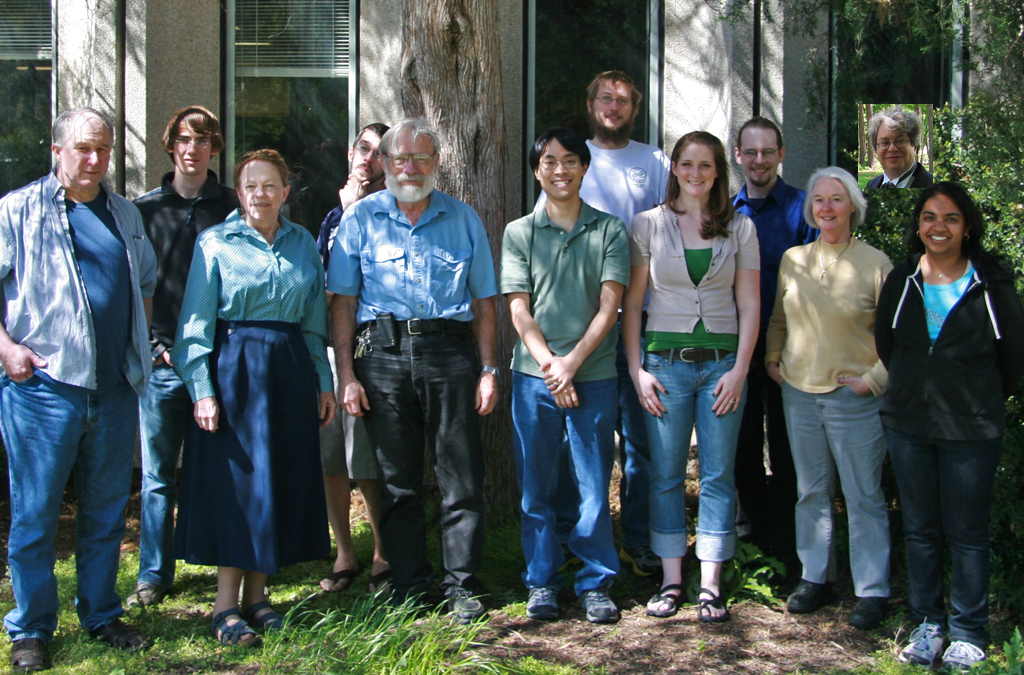

A group photo of the Richardson Lab.
Former Richardson graduate student Bradley Hintze Ph.D. said that Jane Richardson was instrumental in convincing him that graduate school could be possible. He has cerebral palsy, which affects his fine motor coordination and makes some kinds of basic research impossible, such as working with pipettes and glassware. Jane helped convince him that working in crystallography — which has a large computational aspect — would be a good path forward.
“After interviewing at five schools I chose Duke for the explicit reason that I could do my research with Jane,” said Hintze. “I am most grateful for the Richardsons’ ultimate concern — people, human beings. Science is very exciting and can conjure deep passion for methodology and discovery. Unfortunately, the humanity of the researchers, in this case mentees, can get lost in that passion. With the Richardsons, that was never the case; the human aspect of their mentees always took precedence over the science. This cultured trust, and ultimately, I believe, a much deeper passion for the research done in the lab.”
Story by Lindsay Key
Photos by Jared Lazarus
Design by Sam Huntley
Produced by University Communications








